Christian Stippel1, Felix Mujkanovic2, Thomas Leimkühler2, Pedro Hermosilla1
1TU Wien, Austria
2Max-Planck-Institute for Informatics, Germany
SIGGRAPH Asia 2025 (Journal Track)
This is the official codebase for Marching Neurons, a novel method for analytically extracting surfaces from neural implicit representations. Our method leverages the piecewise linear structure of neural networks to traverse the network neuron by neuron, identifying regions that contain the desired surface. The resulting meshes faithfully capture the full geometric information from the network without ad-hoc spatial discretization, achieving unprecedented accuracy across diverse shapes and network architectures while maintaining competitive speed.
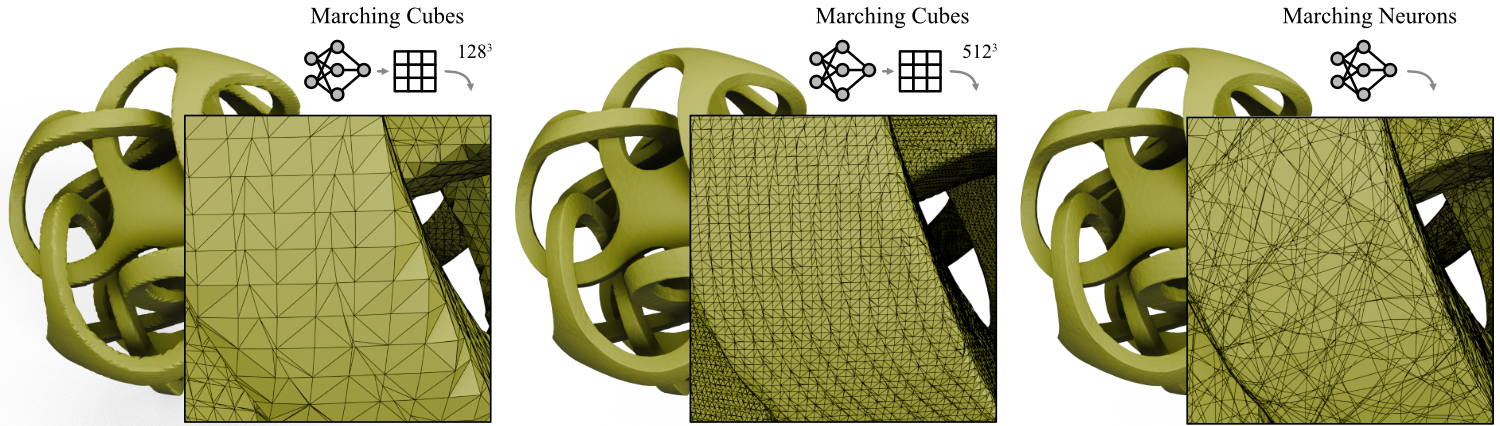
Surfaces extracted from a signed distance function (SDF) represented by a neural network using Marching Cubes with different grid resolutions (left and center) compared to our analytic method (right). While Marching Cubes struggles to reconstruct sharp edges even at high grid resolutions, our analytic method is able to reconstruct the surface accurately.
Download the required data and pretrained models from Google Drive:
Download link: https://drive.google.com/drive/folders/1Jv6J3q8zuoqSGC3WNt_N2rGTIXxBYhOp?usp=drive_link
The folder contains three archives:
nets.zip: Pretrained neural networkszero_level_sets.zip: Ground truth zero-level sets for evaluationmeshes.zip: Sample meshes for training
After downloading, extract all three archives in the project root:
unzip nets.zip # Extracts pretrained networks
unzip zero_level_sets.zip # Extracts ground truth data
unzip meshes.zip # Extracts sample meshes to data/meshes/This will create the following directory structure:
marching_neurons/
├── nets/
│ └── from_sdf/
│ └── {architecture}/
│ └── {mesh_name}/
│ ├── net.pt # Pretrained network
│ └── zero_level_set.ply # Ground truth for evaluation
├── data/
│ └── meshes/ # Sample meshes for training
...
For the complete mesh datasets used in the paper, download and extract them from their respective sources:
- ABC Dataset: https://deep-geometry.github.io/abc-dataset/
- FAUST Dataset: https://faust-leaderboard.is.tuebingen.mpg.de/
- ShapeNet: https://shapenet.org/
- Stanford 3D Scanning Repository: http://graphics.stanford.edu/data/3Dscanrep/
- Thingi10K: https://ten-thousand-models.appspot.com/
Place the downloaded meshes in data/meshes/ and follow the preprocessing steps below.
This project has been tested on:
- OS: Ubuntu 24.04 LTS
- GPU: NVIDIA GPUs with CUDA support
- Python: 3.10 or higher (tested with 3.11)
Create and activate the environment using the provided environment.yml file:
conda env create -f environment.yml
conda activate marching_neuronsThis will install PyTorch and JAX with CUDA 12 support. If you have a different CUDA version, you may need to adjust the installation:
- For CUDA 11.8: After creating the environment, reinstall PyTorch with:
conda activate marching_neurons pip install torch --index-url https://download.pytorch.org/whl/cu118 pip install "jax[cuda11_pip]"
If you prefer to use a Python virtual environment with pip:
# Create a virtual environment (requires Python 3.10 or higher)
python3.11 -m venv venv
# Activate the virtual environment
source venv/bin/activate # On Linux/macOS
# or
venv\Scripts\activate # On Windows
# Install remaining dependencies
pip install -r requirements.txtThe project provides several scripts for the complete pipeline from data preprocessing to surface extraction and evaluation.
Extract surfaces from pretrained networks (requires nets.zip downloaded and extracted):
# Extract triangle mesh using marching neurons (default)
python scripts/extract.py --task from_sdf --arch relu_mlp_d4_w128 --mesh Stanford_bunnyExtracted mesh will be saved as nets/from_sdf/relu_mlp_d4_w128/Stanford_bunny/extract_marching_neurons_triangulated.ply.
Evaluate extracted meshes against ground truth (requires zero_level_sets.zip downloaded and extracted):
python scripts/measure.py --task from_sdf --arch relu_mlp_d4_w128 --mesh Stanford_bunnyTo train your own networks and extract meshes (requires meshes.zip downloaded and extracted):
Follow the complete pipeline below.
Convert ground truth meshes into SDF point clouds for training (requires meshes.zip extracted to data/meshes/):
# Process a single mesh
python scripts/preprocess.py --mesh Stanford_bunny
# Process all meshes in data/meshes/
python scripts/preprocess.pyThis creates SDF point clouds in data/sdf_point_clouds/.
Train neural networks on the preprocessed data:
# Train specific architecture and mesh with a time budget
python scripts/train.py --task from_sdf --arch relu_mlp_d4_w128 --mesh Stanford_bunny --runtime 1:00:00
# Train multiple architectures
python scripts/train.py --task from_sdf --arch relu_mlp_d4_w128 relu_mlp_d4_w256 --mesh Stanford_bunny --runtime 0:30:00Trained networks are saved in nets/{task}/{arch}/{mesh}/net.pt.
Extract surfaces from trained networks using marching neurons:
# Extract triangle mesh (default)
python scripts/extract.py --task from_sdf --arch relu_mlp_d4_w128 --mesh Stanford_bunnyExtracted meshes are saved as nets/{task}/{arch}/{mesh}/extract_marching_neurons_triangulated.ply.
Simplify extracted meshes using QEM (Quadric Error Metric) decimation:
# Simplify meshes to 10% of original triangle count (default)
python scripts/simplify.py --task from_sdf --arch relu_mlp_d4_w128 --mesh Stanford_bunny
# Use a different simplification ratio (e.g., 20%)
python scripts/simplify.py --task from_sdf --arch relu_mlp_d4_w128 --mesh Stanford_bunny --ratio 0.2
# Simplify all meshes in an architecture
python scripts/simplify.py --task from_sdf --arch relu_mlp_d4_w128Simplified meshes are saved as nets/{task}/{arch}/{mesh}/extract_{extractor_name}_qem.ply with statistics in simplify_{extractor_name}_qem.csv.
Compute precision, recall, and triangle count metrics for extracted meshes (requires zero_level_sets.zip downloaded and extracted):
# Measure marching neurons extractions (default: marching_neurons_triangulated)
python scripts/measure.py --task from_sdf --arch relu_mlp_d4_w128 --mesh Stanford_bunny
# Measure all available extractions in a task
python scripts/measure.py --task from_sdfMetrics are saved as nets/{task}/{arch}/{mesh}/metrics_{extractor_name}.csv.
Generate summary statistics, plots, and comparison tables:
# Summarize all results for a task
python scripts/summarize.py --task from_sdf
# Summarize specific architectures
python scripts/summarize.py --task from_sdf --arch relu_mlp_d4_w128 relu_mlp_d4_w256Results are saved in summary/:
summary.csv: Aggregated metricstable.tex: LaTeX table for paper (if using comparison methods)boxplots.pdf: Comparison plots (if using comparison methods)runtime.pdf: Runtime analysis plots (if using comparison methods)
To understand how the core algorithm operates on individual cells in 2d refer to:
python scripts/example_2d.pyIf you find this work useful, please consider citing:
@article{stippel2025neurons,
title={Marching Neurons: Accurate Surface Extraction for Neural Implicit Shapes},
author={Stippel, Christian and Mujkanovic, Felix and Leimk{\"u}hler, Thomas and Hermosilla, Pedro},
journal={ACM Transactions on Graphics (Proc. SIGGRAPH Asia)},
year={2025},
}This work was partially funded by the Austrian Research Promotion Agency (FFG) under the project "AUTARK" (No. 999922723), within the call "Schlüsseltechnologien im produktionsnahen Umfeld". We gratefully acknowledge Wolfgang Koch for the insightful and valuable research discussions.