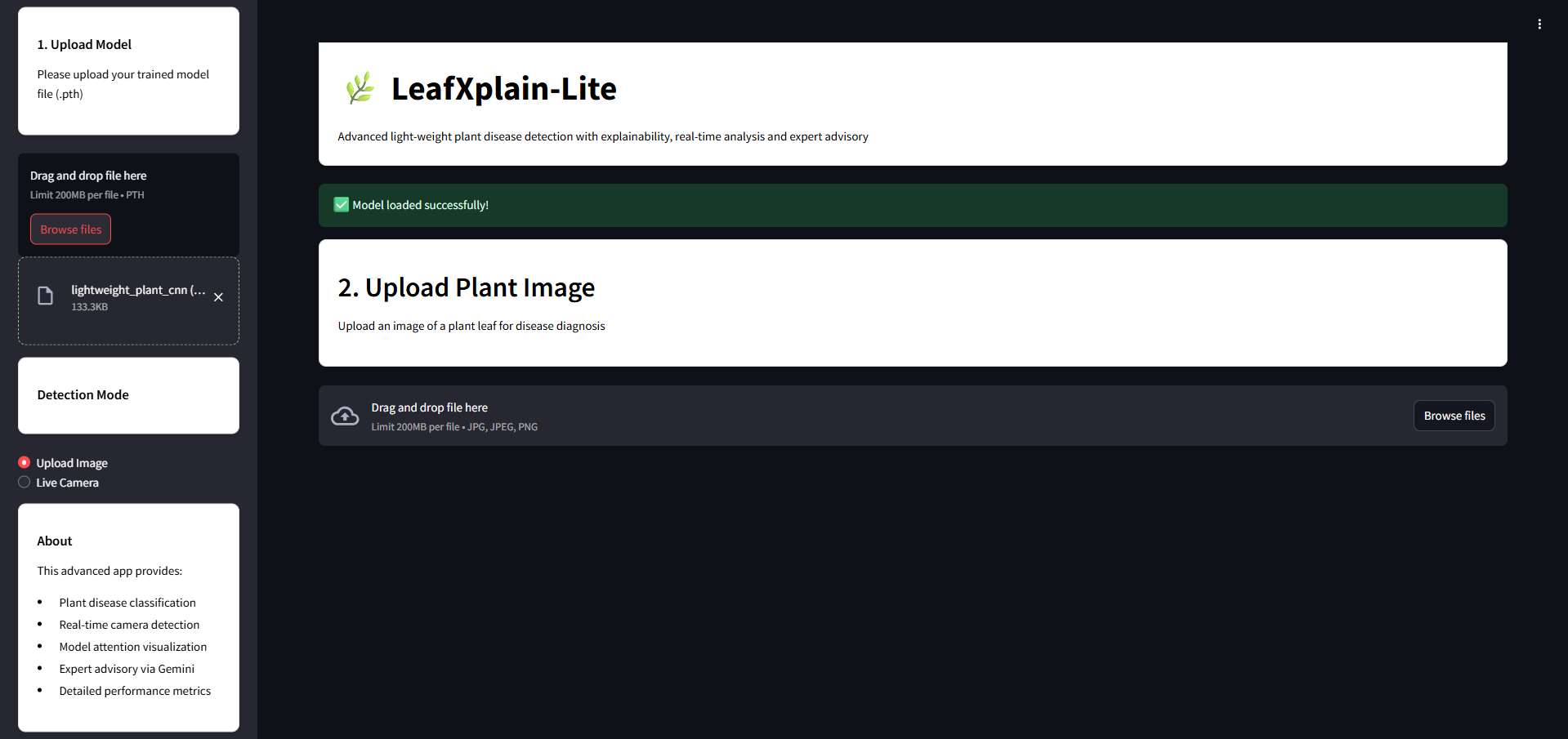
LeafXplain-Lite is a smart, explainable AI-based web application for real-time plant disease detection. It provides intuitive visualizations, interpretable model outputs, and expert treatment suggestions through Gemini AI — all in one lightweight package.
- Upload a leaf image to get a fast and accurate disease diagnosis.
- Powered by a CNN model with attention mechanisms and Grad-CAM explanations.
- Visualizes which part of the leaf influenced the model's decision.
When an image is uploaded, the app provides a quick summary of predictions, including:
- predicted disease
- Grad-Cam Visuals
- Predicted Result
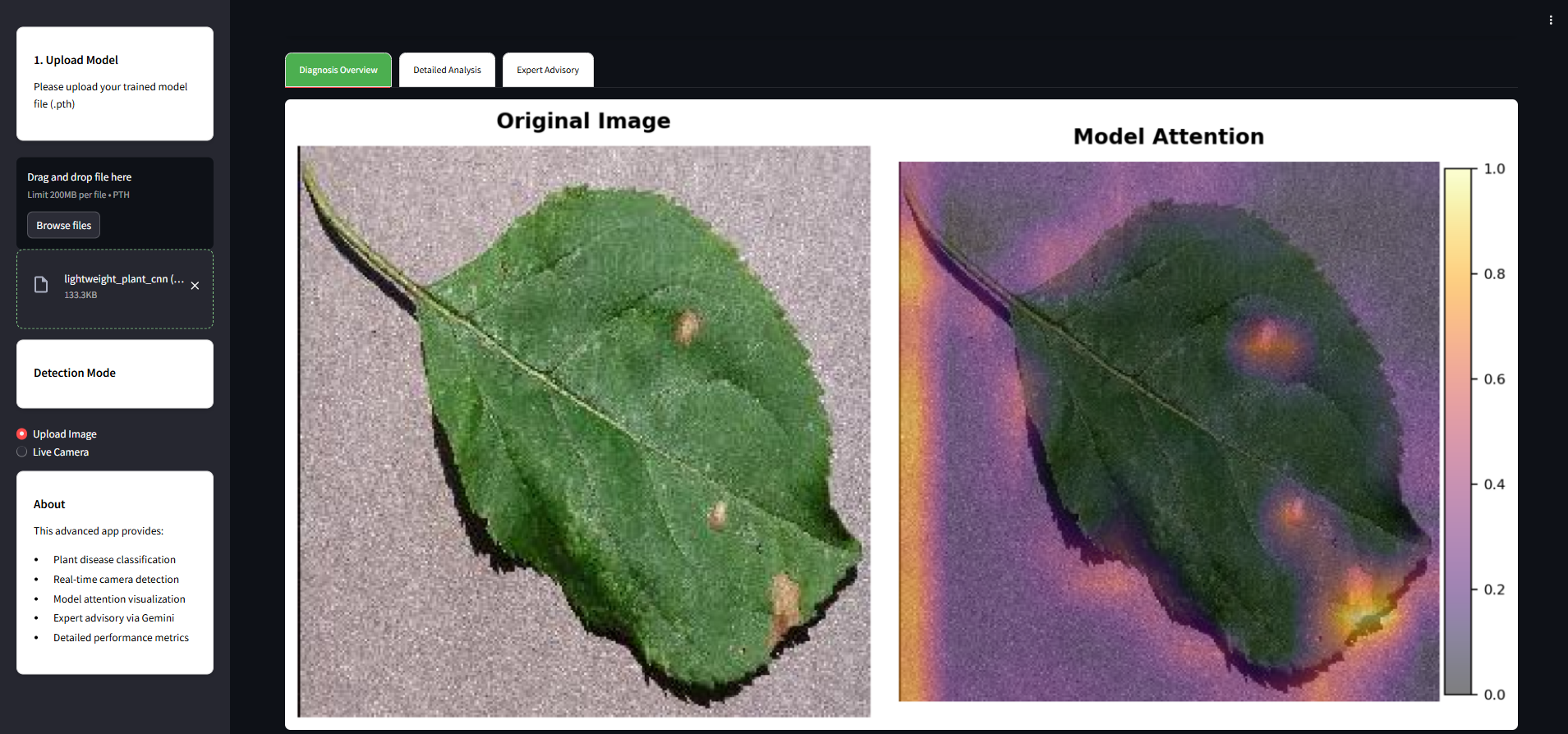
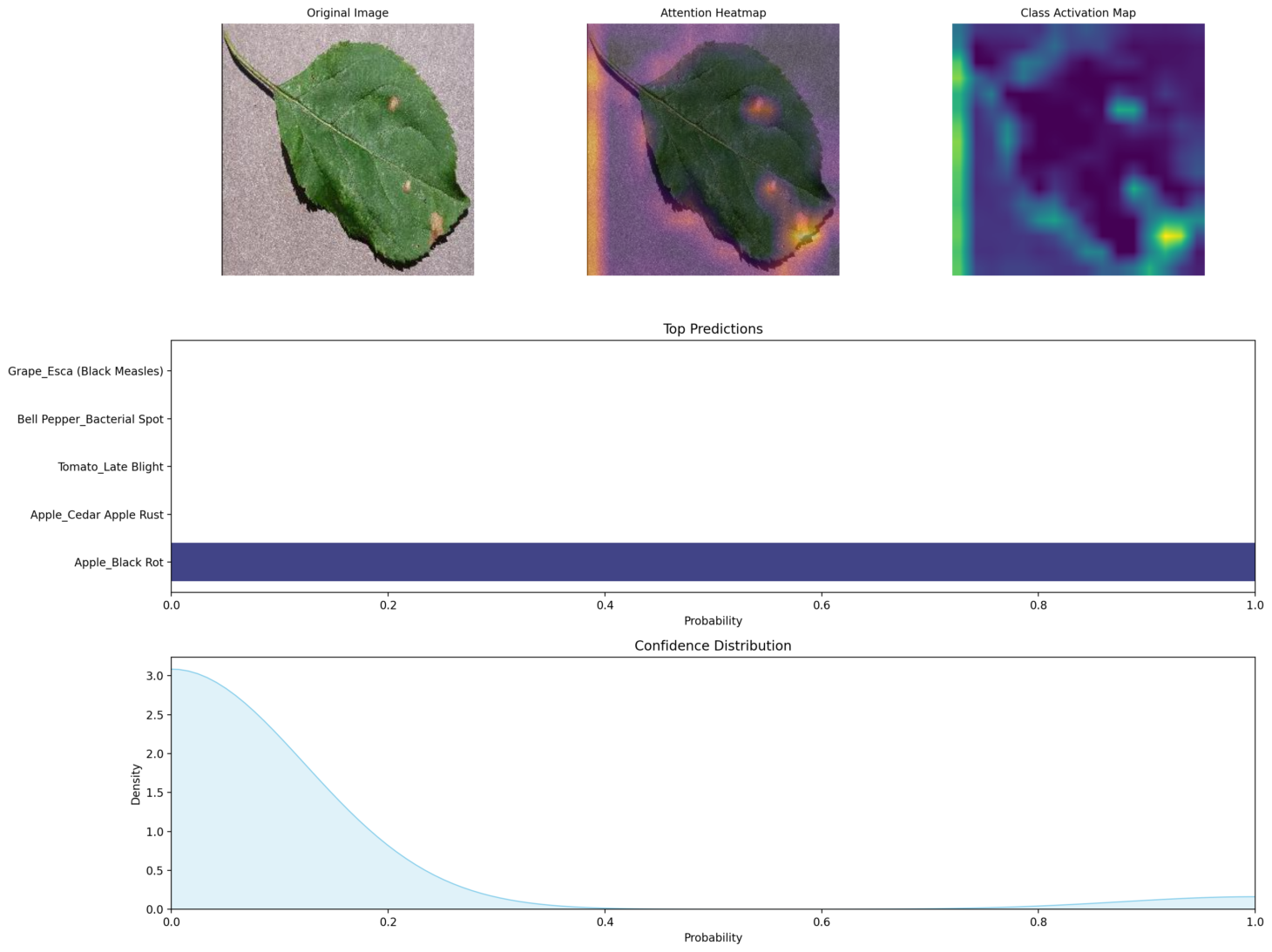
This section breaks down the prediction with clear, scientific visualizations:
- Original Image: What you uploaded.
- Attention Heatmap: Shows what the model is focusing on.
- Class Activation Map (CAM): Highlights influential image regions.
- Prediction Distribution: Probability scores for each class.
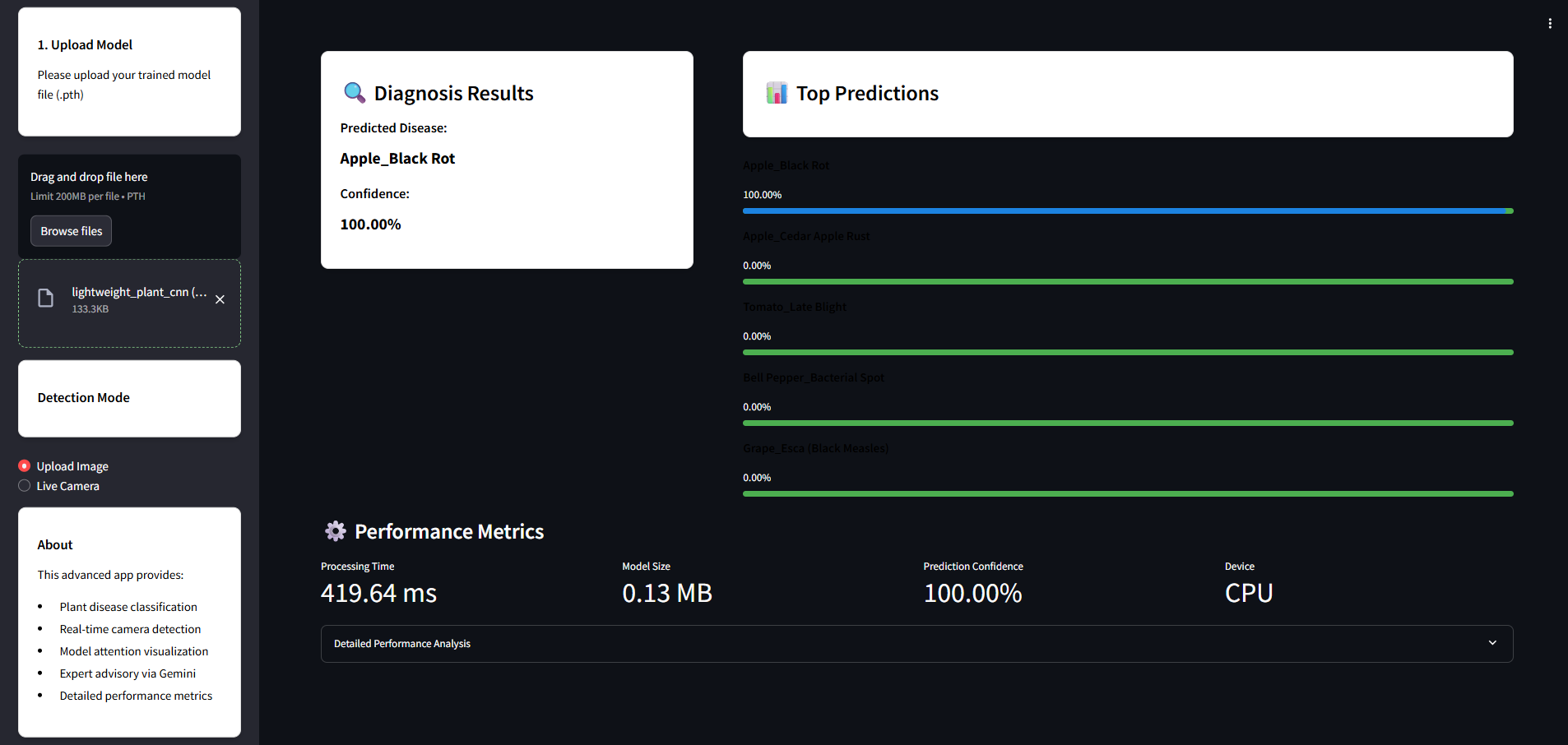
Once a prediction is made, LeafXplain-Lite provides the diagonosis results:
- Predicted Disease
- Disease Confidence
- Top Predictions
- Processing Time
- Model Size
- Switch to Live Camera Mode for real-time disease detection.
- Get instant visual feedback with bounding boxes and overlays.
- Gemini-generated advice is just one click away.
- Python 3.8+
- Streamlit
- PyTorch
- Google Generative AI SDK
git clone https://github.com/AbidHasanRafi/LeafXplain-Lite.git
cd LeafXplain-Lite
pip install -r requirements.txtstreamlit run app.py- Get your API key from Google
- Add to Streamlit secrets or directly in code (for testing)
- Use the default provided model or upload your own
.pth - Model switching is available via dropdown
# model architecture
class PlantDiseaseCNN(nn.Module):
def __init__(self, num_classes):
super(PlantDiseaseCNN, self).__init__()
self.features = nn.Sequential(
nn.Conv2d(3, 32, kernel_size=3, padding=1, bias=False),
nn.BatchNorm2d(32),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2),
nn.Conv2d(32, 32, kernel_size=3, padding=1, groups=32, bias=False),
nn.Conv2d(32, 64, kernel_size=1, bias=False),
nn.BatchNorm2d(64),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2),
nn.Conv2d(64, 64, kernel_size=3, padding=1, groups=64, bias=False),
nn.Conv2d(64, 128, kernel_size=1, bias=False),
nn.BatchNorm2d(128),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2),
nn.Conv2d(128, 128, kernel_size=1),
nn.Sigmoid()
)
self.classifier = nn.Sequential(
nn.AdaptiveAvgPool2d(1),
nn.Flatten(),
nn.Dropout(0.5),
nn.Linear(128, num_classes)
)
def forward(self, x):
features = self.features[:-1](x)
attention = self.features[-1](features)
attended = features * attention
return self.classifier(attended)Supports detection of 20 common plant diseases, including:
- Apple Scab
- Tomato Late Blight
- Grape Black Rot
- Corn Common Rust
- Potato Early Blight ... and more.
- Explainable AI — Grad-CAM shows where the model focuses
- Real-time Detection — From image uploads or live camera
- Gemini-Powered Expertise — Instant, actionable advice
- Lightweight — Small model size, fast on CPU or GPU
- Clean UI — Tab-based view support
- Project Maintainer – Md. Abid Hasan Rafi
- Project Contributor – Pankaj Bhowmik
- GitHub Repository – https://github.com/abidhasanrafi/leafxplain-lite