DiffNMR: Diffusion Models for Nuclear Magnetic Resonance Spectra Elucidation
Qingsong Yang#, Binglan Wu#, **Xuwei Liu, Bo Chen, Wei Li, Gen Long†, Xin Chen†, Mingjun Xiao†
1 Department of Computer Science, University of Science and Technology of China (USTC)
2 Suzhou Laboratory
3 Baidu Inc.
# Equal contribution · † Corresponding authors
DiffNMR is an end-to-end framework that infers molecular structures directly from 1H/13C NMR spectra using a conditional discrete diffusion model. It holistically refines molecular graphs through denoising steps (instead of autoregressive token-by-token generation), improving global consistency and reducing error accumulation. The system couples a two-stage pretraining pipeline—(i) Diffusion Autoencoder (Diff-AE) for molecular representation + graph decoder pretraining and (ii) contrastive alignment between spectra and molecular representations—with a domain-tailored NMR encoder that leverages RBF encodings for chemical shifts and coupling constants. Inference further benefits from similarity-based filtering and optional retrieval-initialized sampling, yielding strong Top‑k accuracy and high Tanimoto similarity across molecule sizes.
- End-to-end spectrum→structure on 1H & 13C NMR.
- Discrete graph diffusion (denoise molecular graphs instead of autoregressive SMILES).
- NMR encoder with RBF embeddings for shifts (and J-coupling) + learnable embeddings for multiplicity & integrals; bi-directional cross‑attention to fuse 1H/13C information.
- Two-stage pretraining: Diff‑AE (molecular encoder + graph diffusion decoder) → contrastive learning to align NMR & molecular spaces.
- Inference enhancements: similarity filtering (cosine scoring in latent space) and retrieval‑initialized sampling.
- MSD multimodal spectroscopic dataset: ~7.9e5 molecules (from USPTO), each with simulated 1H NMR, 13C NMR, HSQC, IR, MS; molecules span 5–35 heavy atoms.
- Evaluation: Top‑k accuracy (exact‑match structure among top candidates) and Tanimoto similarity (Morgan fp, radius 2, 2048 bits).
Observations
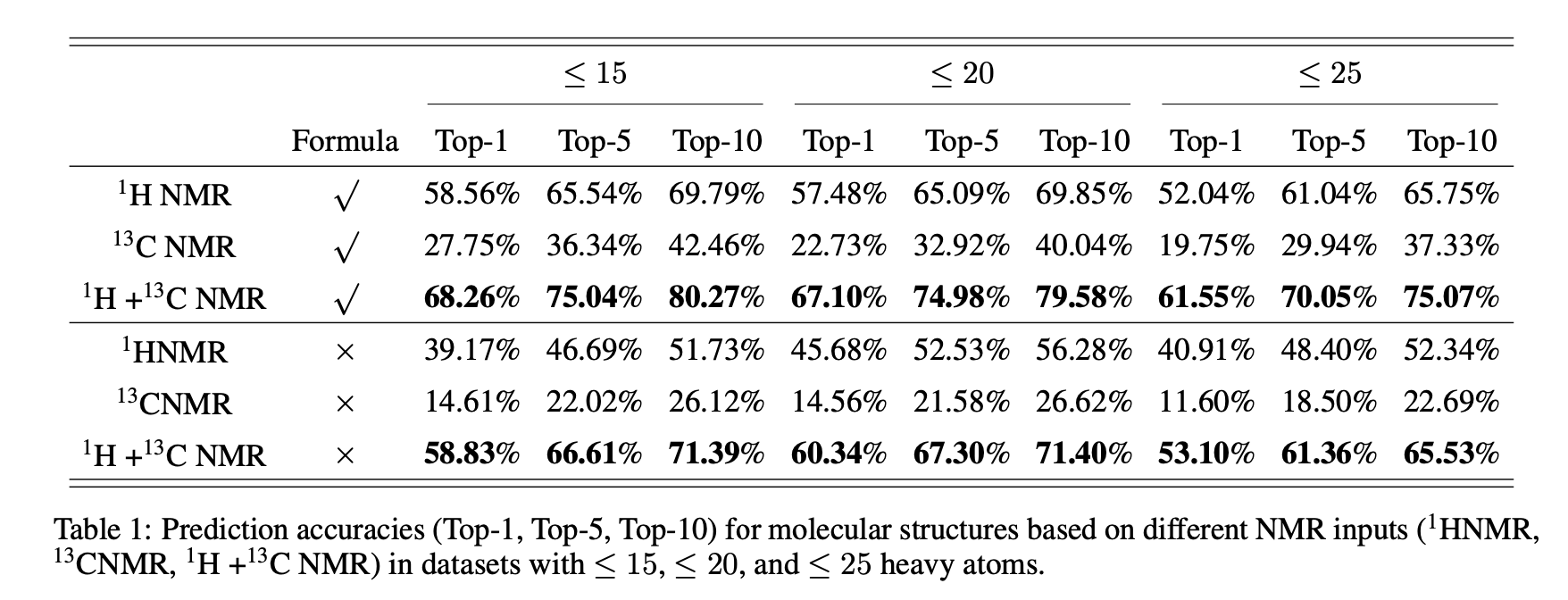
- 1H+13C outperforms single‑modality inputs.
- Providing molecular formula improves accuracy across sizes.
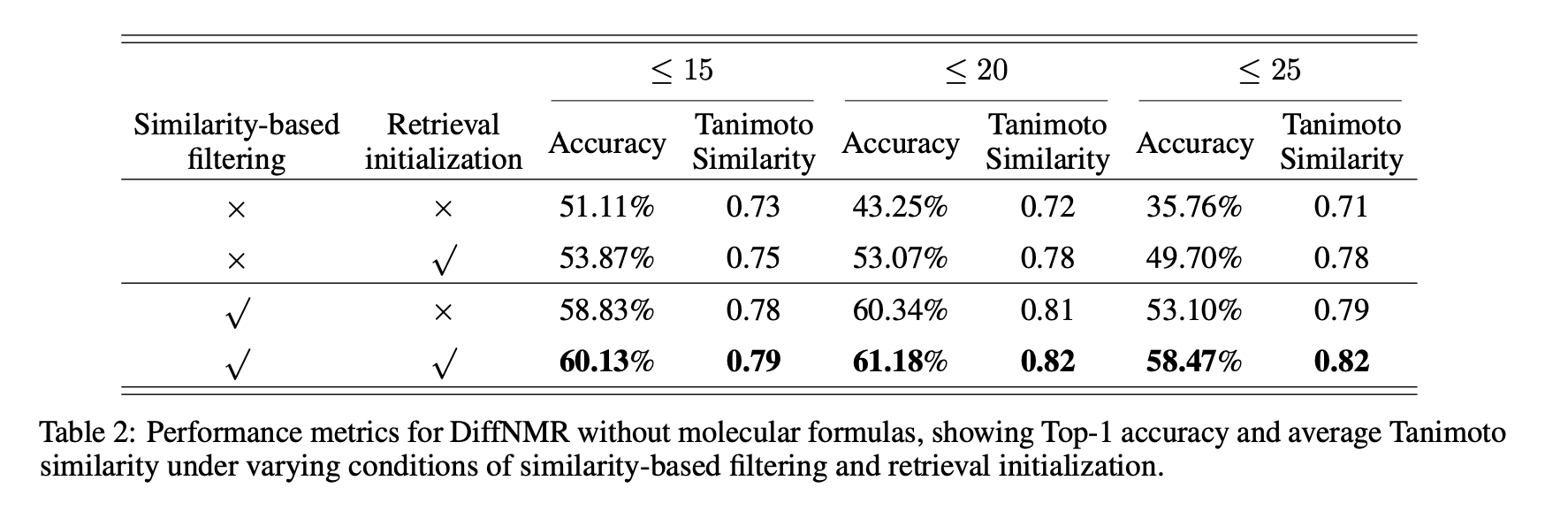
- Similarity filtering and retrieval initialization substantially boost Top‑1 accuracy and average Tanimoto, especially for larger molecules.
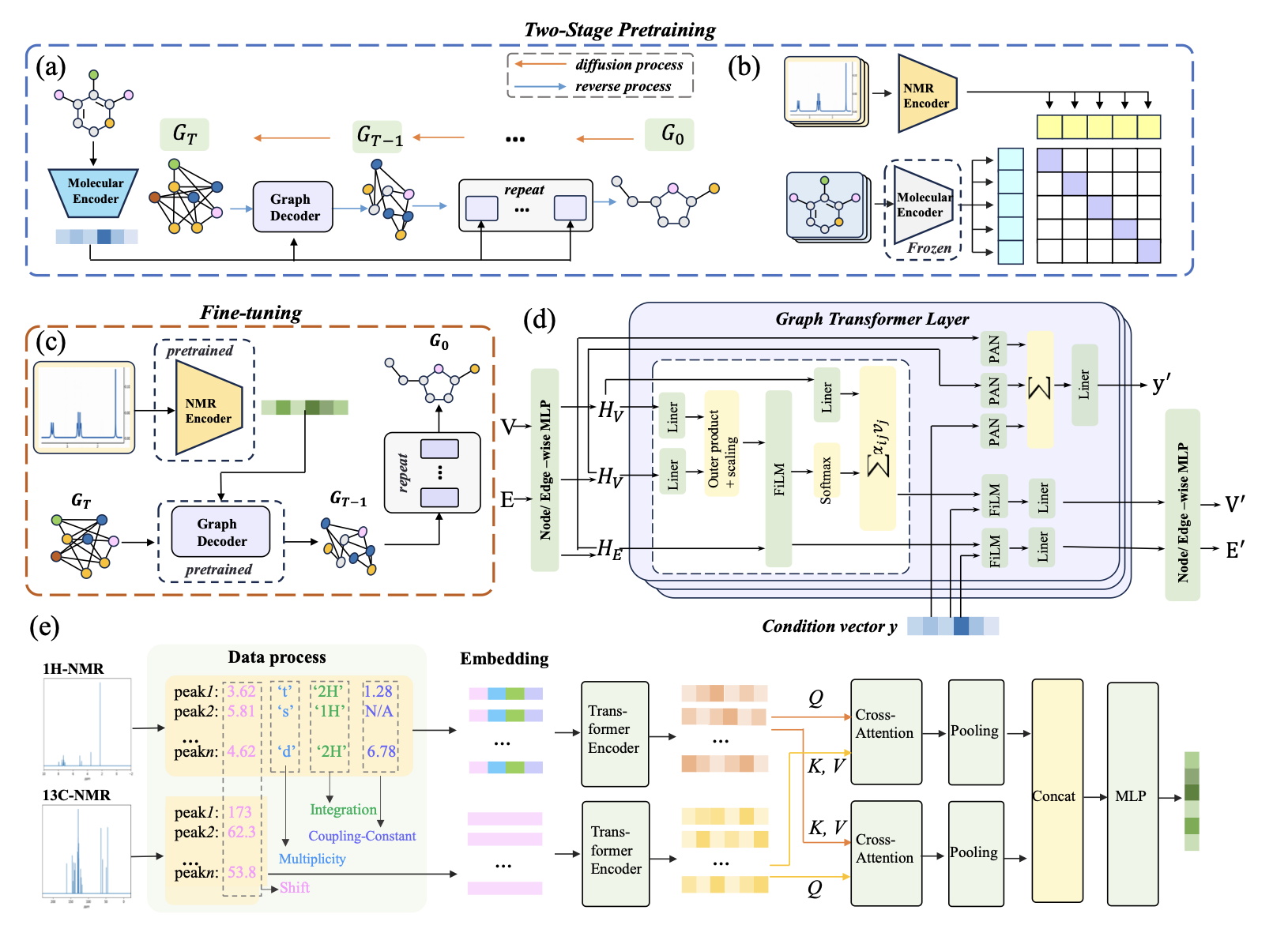
Architecture
- Molecular encoder (graph transformer) → compact molecular representation.
- Graph diffusion decoder (discrete denoising on nodes/edges).
- NMR encoder: RBF(δ) for shifts; embeddings for multiplicity/integrals; RBF(J) for couplings; transformers per modality; bi‑directional cross‑attention (1H↔13C); pooled & fused to a conditioning vector.
Training
- Stage‑1: Diff‑AE — pretrain molecular encoder + diffusion decoder via reconstruction.
- Stage‑2: Contrastive — freeze molecular encoder; train NMR encoder to align to the molecular space with InfoNCE.
- Fine‑tuning — end‑to‑end spectra→graph with diffusion decoder conditioned on NMR embedding.
Inference
- Similarity filtering: rank sampled candidates by cosine similarity between predicted molecule & input NMR embeddings.
- Retrieval initialization (start from nearest neighbor in latent space) improves Top‑1 accuracy.
- Use formula outputs atoms enforced to reset according to formula.
- 1H+13C achieves the best Top‑1/Top‑k; adding molecular formula improves all sizes (≤15/≤20/≤25 HAC).
- Similarity filtering and retrieval initialization notably improve Top‑1 and avg. Tanimoto, with the largest gains on higher HAC.
PaddleMaterials/
spectrum_elucidation/
configs/DiffNMR/
DiffNMR_DiffGraphFormer.yaml
DiffNMR_NMRNet.yaml
DiffNMR.yaml
train.py
sample.py
Refer to the install doc to install PaddleMaterials.
cd spectrum_elucidation
wget https://paddle-org.bj.bcebos.com/paddlematerials/checkpoints/spectrum_elucidation/diffnmr/vocab.tar.gz
tar -xvf vocab.tar.gz
wget https://paddle-org.bj.bcebos.com/paddlematerials/checkpoints/spectrum_elucidation/diffnmr/retrival_database.zip
unzip retrival_database.zipcd pretrained
wget https://paddle-org.bj.bcebos.com/paddlematerials/checkpoints/spectrum_elucidation/diffnmr/DiffNMR_nless15_best.pdparams
cd ..
python spectrum_elucidation/sample.py --config_path='spectrum_elucidation/configs/diffnmr/DiffNMR.yaml' --weights_name='DiffNMR_nless15_best.pdparams' --save_path='result_diffnmr_nless15/' --checkpoint_path="pretrained"revise the spectrum_elucidation/configs/diffnmr/DiffNMR.py and replace the following arguments according to your needs:
flag_retrival_sampling: True
flag_use_formula: True
flag_retrival_initilization: True
num_candidates: 1 # recommend set to 20 for retrieval-sampling similarity filteringrun the command reference to step 2.
If you use DiffNMR, please cite:
@misc{yang2025diffnmr,
title = {DiffNMR: Diffusion Models for Nuclear Magnetic Resonance Spectra Elucidation},
author = {Yang, Qingsong and Wu, Binglan and Liu, Xuwei and Chen, Bo and Li, Wei and Long, Gen and Chen, Xin and Xiao, Mingjun},
year = {2025},
eprint = {2507.08854},
archivePrefix = {arXiv},
primaryClass = {physics.chem-ph},
doi = {10.48550/arXiv.2507.08854},
url = {https://arxiv.org/abs/2507.08854}
}This repository is released under the Apache-2.0 license (unless otherwise stated). See LICENSE for details.
Supported by the National Science and Technology Major Project (2023ZD0120702) and Basic Research Program of Jiangsu (BK20231215). We thank contributors of PaddlePaddle & PaddleMaterials